About ACMG
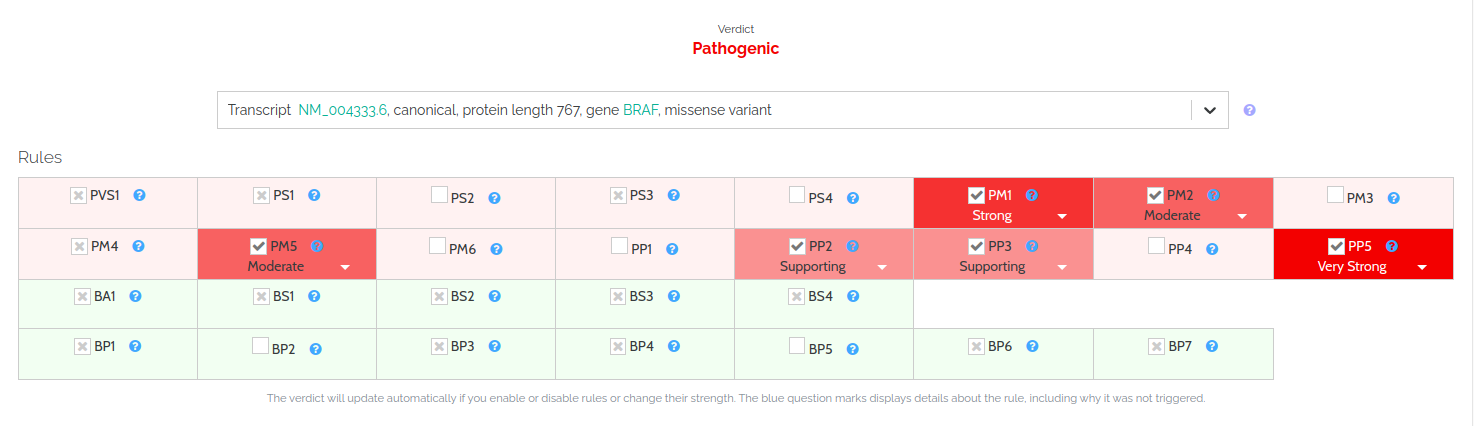
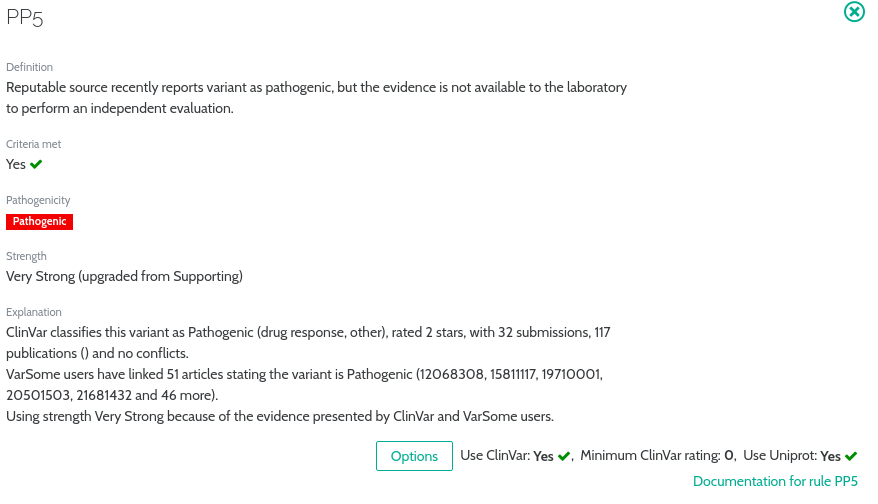
Our implementation is derived from the ”Standards and guidelines for the interpretation of sequence variants” was published in 2015 by Sue Richards et al. in their seminal paper (ACMG Guidelines). Following the advice from our clinical advisors, feedback from the VarSome user community, and using statistically justified thresholds we always strive to provide the most accurate and up-to-date implementation!